NEXTFLEX RNA-Seq 2.0 UDI Barcodes (1-48)
Error-resistant indexing supports cleaner RNA-seq data
Product Description
This NEXTFLEX® RNA-Seq 2.0 Unique Dual Index Barcodes Kit contains 24 uniquely dual- indexed barcoded DNA adapters in plate format for a total of 48 reactions.
The NEXTFLEX™ RNA-Seq 2.0 UDI Barcodes are indexed adapters for single or paired-end sequencing with the NEXTFLEX RNA-Seq 2.0 workflow. Each 8-nt index pair is separated by at least a three-base edit distance to enable error-tolerant demultiplexing and to limit index hopping on patterned flow cells.
Features of RNA-seq 2.0 UDI Barcodes
- UDI full-length adapters for multiplexing up to 384 libraries on Illumina® and Element® AVITI™ platforms
- Tested with the NEXTFLEX RNA-Seq 2.0 kit and compatible with TruSeq-style T-overhang library preps
- Unique dual-index design with ≥3-base edit distance reduces index hopping and sample mis-assignment on patterned flow cells
- Each lot is sequence-verified for index purity by NGS-based QC
- Compatible with NEXTFLEX Universal Blockers
Browse NEXTFLEX RNA-seq 2.0 UDI Barcode options
Error-resistant indexing
The NEXTFLEX RNA-Seq 2.0 UDI Barcodes are built on a dual 8-nt index architecture, with each pair separated by a Hamming distance of three or more bases. This spacing helps prevent single-base substitution errors from producing false index assignments during demultiplexing. In addition, requiring matched i7 and i5 index pairs ensures that reads are only assigned when both ends correspond to the correct sample. This dramatically reduces the risk of misassignment from index hopping or low-level contamination, issues that are especially problematic on patterned flow cells.
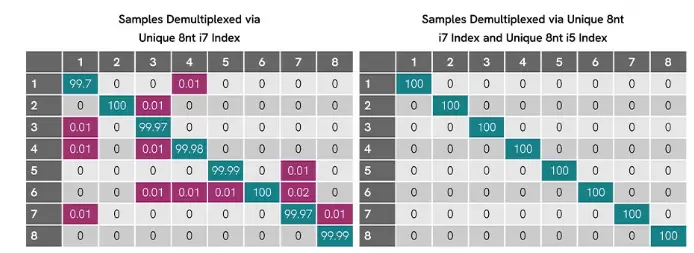
Figure 1. Dual-index design eliminates misassignment on Illumina® MiSeq®. Eight libraries prepared with NEXTFLEX Barcodes show low off-target assignment (≤0.02%) when demultiplexed using i7-only indices (left). When demultiplexed with matched i7 + i5 index pairs (right), all reads are correctly assigned with no detectable cross-contamination.
These design features contribute directly to data quality. Researchers retain more usable reads, avoid index collisions, and maintain better sample balance across RNA-seq libraries. The result is greater confidence in read-level assignments for applications such as differential expression, isoform detection, and low-input RNA profiling, where even low rates of cross-sample contamination can compromise biological interpretation.
Seamless workflow integration
NEXTFLEX RNA-Seq 2.0 UDI Barcodes are compatible with any TruSeq-style T-overhang ligation workflow, including PCR-free and amplified RNA-seq protocols. The plate-based format supports flexible library prep planning, whether you're running high-throughput batches or low-plex experiments, while preserving index integrity across stored wells.
Quality-controlled barcodes for trusted results
Each index set undergoes sequencing-based quality control to confirm purity and assignment accuracy. Proprietary contamination reduction methods lower cross-sample crosstalk to ~ 0.1%. Color balancing and avoidance of long homopolymers further support even signal distribution and reduce sequencing noise. Controlled manufacturing ensures traceable, lot-to-lot consistency for reproducible results across runs.
Scalability, blocker compatibility, and UMI options
NEXTFLEX RNA-Seq 2.0 indexing scales from 24 or 96 UDI pairs up to 384 unique combinations across four plates, without changing demultiplexing settings or workflow parameters. For added hybridization specificity, NEXTFLEX Universal Blockers can be used to improve on-target rates. For experiments requiring molecule-level quantitation, NEXTFLEX UDI-UMI Adapters incorporate UMIs directly into the indexing structure to support duplicate collapsing and accurate counting.
- Catalog Number
NOVA-512920-EVAL48-REV - Supplier
Revvity - Size
- Shipping
Dry Ice

