QuantiMir Kit
| Specifications | |
|---|---|
| Organism: | Human, Mouse |
Product Description
- Simple and robust procedure for converting small RNAs from any tissue source into ready-to-use cDNA templates for real-time qPCR
- Validate newly discovered microRNAs by quantifying them
- Quanititate microRNA expression from any tissue source
- Detect and quantitate siRNAs, shRNAs, and mRNAs in knockdown experiments
- No more tedious Northerns or RNase protection assays
- Stop wasting your precious RNA on single microRNA detection per RT reaction
- Convert enough RNA to cDNA for up to 5,000 qPCR measurements
- Amenable to high-throughput screening of clinical tissue samples including laser-captured micro-dissected specimens
- Entire procedure takes <2 hours
- Three reference assays included in each kit: Human U6, Mouse U6 and Human/Mouse miR-16
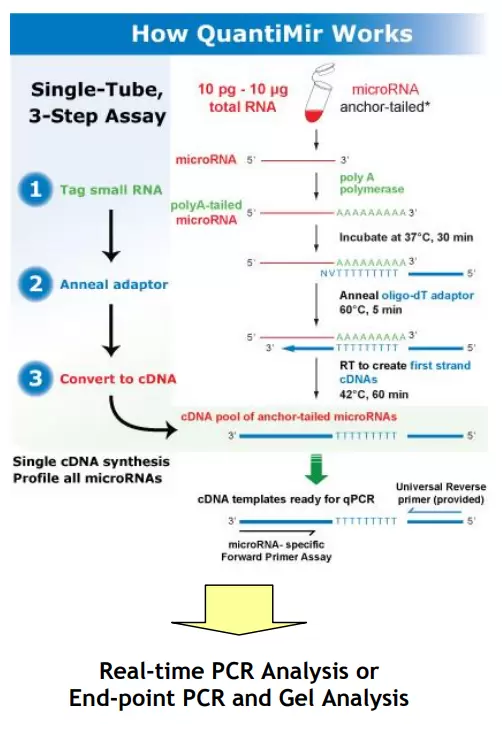
The QuantiMir™ RT Kit from SBI is designed to anchor-tail and convert all small, non-coding RNAs into cDNA starting from total RNA samples. First a poly-A tail is added to the 3‘ end of small non-coding RNAs in the presence of poly-A polymerase. An oligo-dT adaptor primer is used to initiate the subsequent reverse transcription reaction. The resulting cDNAs are ready-to-use for either end-point PCR experiments or to perform real-time qPCR analysis.

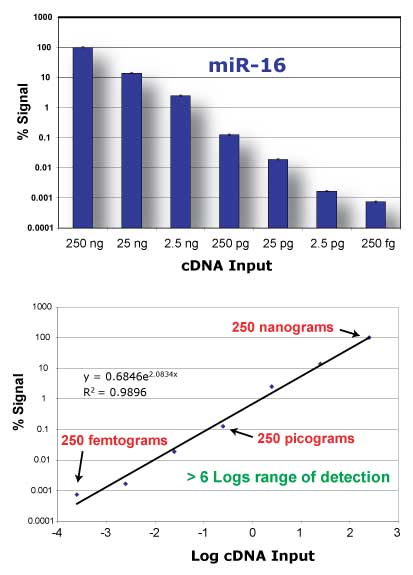
Sensitivity
You can start with picograms to micrograms of Total RNA, even with fractionated small RNA samples and can detect microRNA expression changes in a dynamic range of at least 6 log-fold differences.
Accuracy
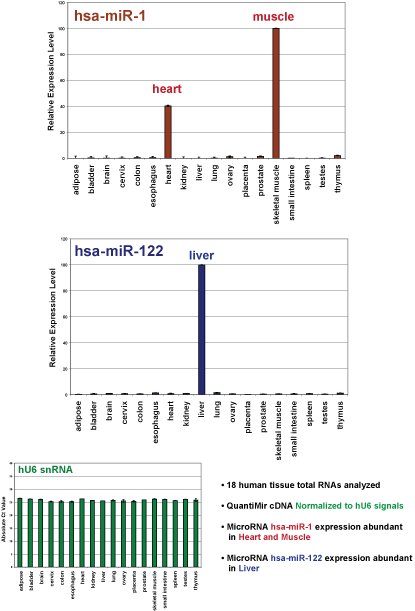
First strand cDNAs were synthesized using 500 ng total RNA from 18 different human tissues: adipose, bladder, brain, cervix, colon, esophagus, heart, kidney, liver, lung, ovary, placenta, prostate, skeletal muscle, small intestine, spleen, testes, and thymus.
The QuantiMir cDNAs from the 18 human tissues were balanced to yield equal Ct values for the U6 snRNA normalizing transcript (Below in Green bars). Real-time PCR results demonstrated this snRNA is uniformly expressed across the 18 tissues examined.
Assays specific for microRNA miR-1 demonstrate specific Heart and Skeletal expression (Below in Red bars) and assays for microRNA miR-122 clearly show specific Liver expression (Below in Blue bars).
Application Examples
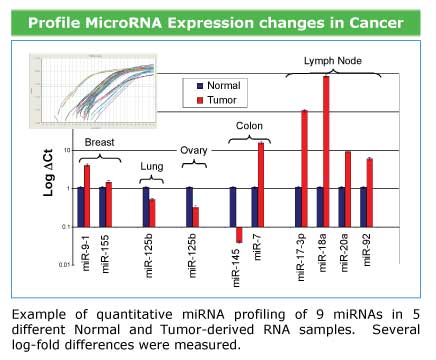
Profile microRNA expression in cancer and normal tissue samples
Numerous miRNAs has been found to have links with some types of cancer. Expression profiling of microRNAs can assist in identifying the type and severity of the oncogenic state and microRNA expression "signatures" can be used to develop diagnostics and therapeutics.
Measure both shRNA expression and mRNA knockdown in a single QuantiMir cDNA sample
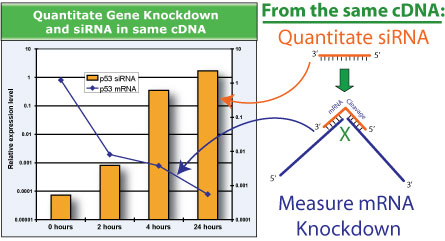
Quantitate siRNAs and mRNA knockdown using QuantiMir: The use of sh/siRNAs to lower the levels of endogenous messenger RNA expression is a powerful and effective tool to study the roles of a targeted transcript in biological systems. Real-time qPCR has become the gold standard in accurately measuring the knockdown effects of specific sh/siRNAs in these experiments. The means of assessing the expression levels of sh/siRNA input has been typically achieved through laborious and nonquantitative Northern blot analyses. Here we present a new method to quantitate both the expression levels of sh/siRNA and the knockdown effect on the targeted messenger RNA transcript using a passive lysis buffer coupled with the QuantiMir kit cDNA synthesis and Real-time qPCR.
After transfection of siRNA or shRNA expression constructs, the cells are removed from the transfection plate using standard trypsinization. The cells are pelleted and subsequently lysed using 100 ul of SBI’s Cells-to-Cts passive lysis Buffer system, which is available in the Cells-to-Cts QuantiMir kit (see link below): A 5 ul aliquot of the lysed cell suspension is then input directly into the QuantiMir RT reaction. The unique technologies in the QuantiMir RT kit enables the conversion of small RNA into detectable cDNA while simultaneously converting messenger RNA into cDNA. Accurate measurement of sh/siRNAs and the messenger RNA transcript in knockdown experiments can be accomplished using the cDNA created from this single RT reaction. The QuantiMir RT reaction generates enough cDNA suitable for numerous qPCR reactions. Shown below is an example of a timecourse study using anti-p53 shRNA-directed knockdown of the endogenous p53 mRNA transcript. From the same QuantiMir cDNA, both the p53 siRNA (Orange bars) and the p53 mRNA transcript (Blue line) were measured by qPCR.
References
- Fortner, D.M. et al. (1994) A stem/loop in U6 RNA defines a conformational switch required for pre-mRNA splicing. Genes and Development 8: 221-233.
- Lee NS, Kim JS, Cho WJ, Lee MR, Steiner R, Gompers A, Ling D, Zhang J, Strom P, Behlke M, Moon SH, Salvaterra PM, Jove R, Kim KS. miR-302b maintains ""stemness"" of human embryonal carcinoma cells by post-transcriptional regulation of Cyclin D2 expression.Biochem Biophys Res Commun. 2008 Dec 12;377(2):434-40.
- Lo HL, Chang T, Yam P, Marcovecchio PM, Li S, Zaia JA, Yee JK. Inhibition of HIV-1 replication with designed miRNAs expressed from RNA polymerase II promoters.Gene Ther. 2007 Sep 6
- Paul E. Neiman, Katrina Elsaesser, Gilbert Loring and Robert Kimmel. Myc Oncogene-Induced Genomic Instability: DNA Palindromes in Bursal Lymphomagenesis. PLoS Genet. 2008 July; 4(7): e1000132.
- Shi, R. and Chiang, V.L. (2005) Facile means for quantifying microRNA expression by real-time PCR. BioTechniques 39:519-525.
- Catalog Number
RA420A-1-SBI - Supplier
SBI System Biosciences - Size
- Shipping
Blue Ice

