Cytoplasmic and Nuclear RNA Purification Kit
| Specifications | |
|---|---|
| Product Category: | Total RNA incl. microRNA Purification |
| Sample Type: | Cultured animal cells, tissue samples |
Product Description
- Fast and easy processing - Rapid spin-column format allows for the processing of 10 samples in 45 minutes.
- No genomic DNA contamination in cytoplasmic fraction - Purified cytoplasmic RNA can be used directly in RT-PCR reactions with no DNase treatment required.
- No phenol or chloroform extractions
- Isolate a diversity of RNA species - All nuclear and cytoplasmic RNA species, from large mRNA species down to microRNA (miRNA) and small interfering RNA (siRNA) can be isolated in high quality.
- This kit is available in both 50 prep and 100 prep sizes
Norgen’s Cytoplasmic & Nuclear RNA Purification Kit provides a rapid method for the isolation and purification of both cytoplasmic and nuclear RNA from cultured animal cells and small tissue samples. In certain circumstances it is desirable to be able to isolate fractionated RNA as opposed to total RNA. For example, it may be preferable to isolate only mature, cytoplasmic RNA for some studies on expression profiling. Alternatively it may be desirable to isolate nuclear RNA in order to investigate and study pre-processed (non-spliced) RNA. Furthermore, this kit can be used to isolate RNA for downstream applications where it is necessary to avoid DNA contamination, since the cytoplasmic fraction has been shown to be free of all traces of genomic DNA. Norgen’s Cytoplasmic & Nuclear RNA Purification Kit can be used to isolate all sizes of RNA from the cytoplasmic and nuclear RNA fractions, including all small RNA species. The kit is supplied with sufficient reagents to perform either 50 cytoplasmic RNA preparations or 25 cytoplasmic and 25 nuclear RNA preparations. The isolated high quality RNA is suitable for use in downstream applications including real-time PCR, reverse transcription PCR, Northern blotting, RNase protection and primer extension, and expression array analysis.
Workflow
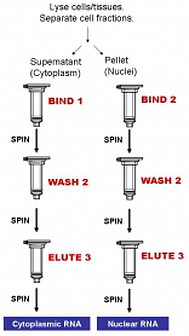
Norgen's Purification Technology
Purification is based on spin column chromatography using Norgen's proprietary resin as the separation matrix. The cytoplasmic RNA is preferentially purified from the nuclear RNA and other cellular components such as proteins, without the use of phenol or chloroform. The process involves first lysing the cells or tissue of interest with the provided Lysis Solution. The lysate is then separated through centrifugation, with the supernatant containing the cytoplasmic RNA and the pellet containing the nuclear RNA. Binding solution and ethanol is then added to the desired fraction, and the solution is loaded onto a spin-column. Norgen's resin binds RNA in a manner that depends on ionic concentrations. Thus only the RNA will bind to the column, while the contaminating proteins and genomic DNA will be removed in the
flowthrough or retained on the top of the resin. The bound RNA is then washed twice with the provided wash buffer in order to remove any remaining impurities, and the purified RNA is eluted with the elution buffer. The purified RNA is of the highest integrity, and can be used in a number of downstream applications including real time PCR, reverse transcription PCR, Northern blotting, RNase protection and primer extension, and expression array assays.
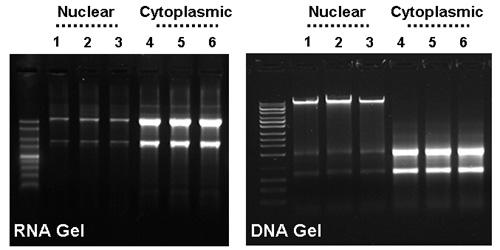
Effective Separation of HeLa Cell Cytoplasmic & Nuclear RNA. Norgen’s Cytoplasmic & Nuclear RNA Purification Kit was used to effectively separate cytoplasmic and nuclear RNA from 0.75 million HeLa cells in triplicate. Panel A: High quality cytoplasmic and nuclear RNA was purified using Norgen's kit. Note the integrity and quality of both cytoplasmic and nuclear RNA. Ten microliters of each 50 µL elution were run on a 1.5% formaldehyde-agarose gel. Lane M is Norgen's 1 kb RNA Ladder, lanes 1-3 contain nuclear RNA and lanes 4-6 contain cytoplasmic RNA. Panel B: Ten microliters of the above cytoplasmic and nuclear RNA isolated from HeLa cells using Norgen's kit was run on a 0.9% agarose DNA gel. Genomic DNA is clearly visible in the nuclear RNA fractions (lanes 1-3), however, no genomic DNA can be detected in the cytoplasmic RNA fractions (lanes 4-6). Note that an optional on-column DNase treatment protocol is provided to remove the genomic DNA in the nuclear fraction.
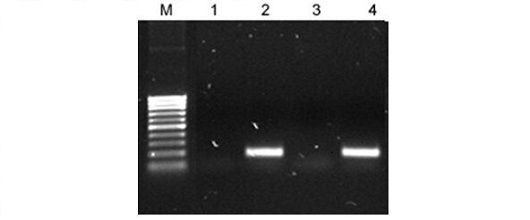

Genomic DNA-free Cytoplasmic RNA. RT-PCR was performed using human beta actin primers on 2 µL of the 50 µL of cytoplasmic RNA isolated from 1 million HeLa cells using Norgen's Cytoplasmic & Nuclear RNA Purification Kit. Lane M is Norgen’s PCR Sizer DNA ladder, Lanes 1 and 3 are the negative control (PCR only, without reverse transcript), and Lanes 2 and 4 are the actual RT-PCR that show the expected 166 bp RT-PCR product. The expected amplicon size from the gene copy is the same as that from the RNA transcript. The lack of product in lanes 1 and 3 indicates that no genomic DNA contamination is present in the isolated cytoplasmic RNA. All PCR products were resolved on a 1X TAE, 2% agarose gel.
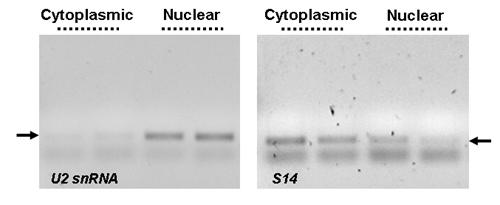
Specific Separation and Subsequent Amplification of Cytoplasmic and Nuclear Transcripts. Norgen’s Cytoplasmic & Nuclear RNA Purification Kit effectively separates cytoplasmic and nuclear RNA. RT-PCR was performed using human U2 snRNA (left panel) or S14 (right panel) primers on 0.5 µg of either cytoplasmic or nuclear RNA fractions isolated from 1 million HeLa cells using Norgen's Cytoplasmic & Nuclear RNA Purification Kit. In the left panel, an intense PCR product was observed only in the nuclear fraction when the nuclear-specific U2 snRNA primers were used. In the right panel, the majority of the PCR product for the house-keeping transcript of S14 was observed in the cytoplasmic fraction. The two results combined showed effective separation of the cytoplasmic and nuclear RNA using Norgen’s Cytoplasmic & Nuclear RNA Purification Kit. All PCR products were resolved on a 1X TAE, 1.5% agarose DNA gel.
Product Citations
- Bin ZhuID, Linhong Li, Rui Wei, Pei Liang*, Xiwu GaoI (2021) Regulation of GSTu1-mediated insecticide resistance in Plutella xylostella by miRNA and lncRNA PLOS Genetics
- Chen J, Zhang K, Zhi Y, et al. (2021) Tumor‐derived exosomal miR‐19b‐3p facilitates M2 macrophage polarization and exosomal LINC00273 secretion to promote lung adenocarcinoma metastasis via Hippo pathway Clinical and Translational Medicine.
- Cheng YQ, Wu CR, Du MR, et al. (2021) CircLphn3 protects the blood-brain barrier in traumatic brain injury Neural Regen Res.
- García‐Padilla C, Domínguez JN, Lodde V, et al. (2021) Identification of atrial‐enriched lncRNA Walras linked to cardiomyocyte cytoarchitecture and atrial fibrillation The FASEB Journal.
- Guo H, Lin S, Gan Z, Xie J, Zhou J, Hu M. (2021) lncRNA FOXD3‑AS1 promotes the progression of non‑small cell lung cancer by regulating the miR‑135a‑5p/CDK6 axis Oncol Lett.
You save 10 %
436,00 €
