Stool Total RNA Purification Kit
| Specifications | |
|---|---|
| Product Category: | Total RNA incl. microRNA Purification |
| Sample Type: | Stool |
Product Description
- Simultaneous isolation of both host RNA and microbial RNA (universal protocol)
- Eliminates PCR inhibitors including humic acid
- Fast and easy processing - Rapid isolation of high-quality, ready-to-use RNA in 30 minutes.
- No organic extraction or alcohol precipitation - Phenol is not required for the isolation
- High yields - Consistent, high yields of inhibitor-free RNA
Norgen’s Stool Total RNA Purification Kit provides a convenient and rapid method to purify total RNA from small amounts of stool samples. All types of stool samples can be processed with this kit, including animal fecal samples and manure. The kit removes all traces of humic acid using the provided Bead Tubes and a combination of chemical and physical homogenization and lysis. A simple and rapid spin column procedure is then used to further purify the RNA. The kit purifies all sizes of RNA, from large mRNA and ribosomal RNA down to microRNA and small interfering RNA. The protocol does not rely on the use of phenol or chloroform, thereby providing a user friendly procedure and allowing high-throughput analysis on the lab bench. The purified RNA is of the highest integrity, and can be used in a number of downstream applications including real time PCR and reverse transcription PCR for gene expression analysis.
The purified RNA is of the highest integrity, and can be used in a number of downstream applications including real time PCR and reverse transcription PCR for gene expression analysis.
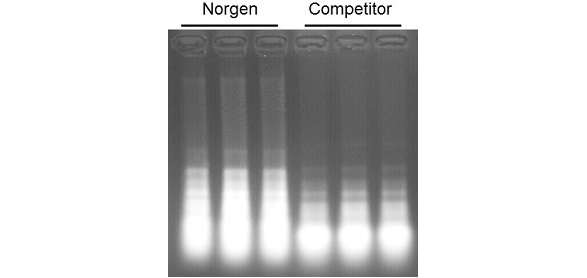
High Yields of Stool RNA Total RNA was purified from 200 mg of human stool using Norgen’s Stool RNA Purification Kit and a leading competitor’s kit, in triplicate. For analysis, 7.5 µL of each 75 µL elution was loaded on a 1.2 % 1x MOPS formaldehyde-agarose gel. Norgen’s kit was found to have a higher yield of RNA, isolated from 200mg of female human toddler stool.
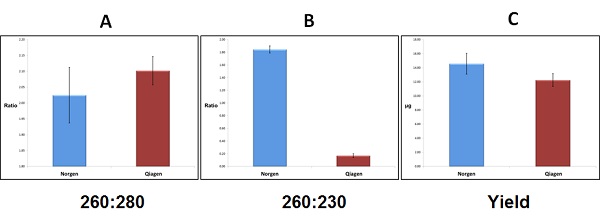
Yield and Quality of Purified Stool RNA. Total RNA was isolated from 200 mg of human stool using Norgen’s Stool RNA Purification Kit and a leading competitor kit. Comparisons were then made based on yield, and A260:A280/A260:A230 ratios measured using the NanoVue Plus™. In Panel A it can be seen that both kits isolated RNA with high A260:A280 ratios (all samples were found to be above 1.8 and below 2.2). In Panel B, Norgen’s kit was found to isolate RNA with a high A260:A230 (with all samples once again falling in the 1.8-2.2 range). The competitor kit, however, was found to isolate RNA with extremely low A260:A230 ratios, with none of the samples displaying a A260:A230 ratio higher than 0.20. The resulst in Panel C are in agreement with the gel photo from Figure 1, and Norgen’s kit was found to isolate higher amounts of RNA, with an average yield of 14.58 µg, compared to the competitor’s average of 12.26 µg.
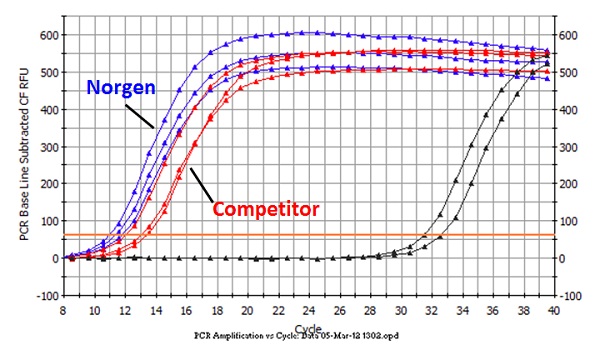
Detection of Bacterial Stool RNA using 16S Primers. Total stool RNA was isolated from 200 mg human stool samples using Norgen’s Stool RNA Purification Kit and a leading competitor’s kit. Five microliters of purified RNA was used in a 20 µL reverse-transcription reaction using Invitrogen’s Superscript III system with 16S reverse primers. The cDNA generated was then used in a qPCR reaction involving Norgen’s 2X PCR Mastermix spiked with SYBR green (Bio-Rad), using 0.3µM of primers against bacterial 16S. As can be seen in the amplification plot, Norgen’s kit outperformed the leading competitor’s kit by on average 1.5 Ct values. This indicates that Norgen isolated higher quality and yields of RNA from stool, that can be used in an array of downstream applications.
Product Citations
- Francavilla A, Gagliardi A, Piaggeschi G, et al. (2021) Faecal miRNA profiles associated with age, sex, BMI, and lifestyle habits in healthy individuals Sci Rep.
- Pisano, A., Griñan-Lison, C., Farace, C., Fiorito, G., Fenu, G., Jiménez, G., Scognamillo, F., Peña-Martin, J., Naccarati, A., Pröll, J., Atzmüller, S., Pardini, B., Attene, F., Ibba, G., Solinas, M. G., Bernhard, D., Marchal, J. A., & Madeddu, R. (2020) The Inhibitory Role of miR-486-5p on CSC Phenotype Has Diagnostic and Prognostic Potential in Colorectal Cancer Cancers
- N Nakayama, M Hamaguchi (2016) Multiplex reverse transcription quantitative PCR detection of a single-stranded RNA virus HcRNAV infecting the bloom-forming dinoflagellate Heterocapsa circularisquama Limnology and Oceanography
- Lopez CC, Serio A, Rossi C, Mazzarrino G, Marchetti S, Castellani F, Grotta L, Fiorentino FP, Paparella A, Martino G (2016) Effect of diet supplementation with Ascophyllum nodosum on cow milk composition and microbiota Journal of Dairy Science
- Maria De Angelis, Eustacchio Montemurno, Lucia Vannini, Carmela Cosola, Noemi Cavallo, Giorgia Gozzi, Valentina Maranzano, Raffaella Di Cagno, Marco Gobbettia and Loreto Gesualdo (2015) The role of whole-grain barley on human fecal microbiota and metabolome Applied and Environmental Microbiology
- Catalog Number
49500-NB - Supplier
Norgen Biotek - Size
- Shipping
RT
You save 10 %
544,00 €
