Soil Total RNA Purification Kit
| Specifications | |
|---|---|
| Product Category: | Total RNA incl. microRNA Purification |
| Sample Type: | Soil |
Product Description
- Isolate high quality total RNA from a variety of soil samples
- Process all types of soil, including common soil, compost and manure
- Remove all traces of humic acids and other inhibitors of PCR
- Isolates all sizes of RNA, including microRNA, without phenol
- Complete kit including bead tubes and humic acid removal columns (HAR)
- Purification is based on spin column chromatography that uses Norgen’s proprietary Silicon Carbide resin separation matrix
Norgen’s Soil Total RNA Purification Kit provides a convenient and rapid method to purify total RNA from small amounts of soil samples. All types of soil samples can be processed with this kit, including common soil samples and difficult soil samples with high humic acid content such as compost and manure. The kit removes all traces of humic acid using the provided Bead Tubes and a combination of chemical and physical homogenization and lysis. A simple and rapid spin column procedure is then used to further purify the RNA.
Purification is based on spin column chromatography using Norgen's resin as the separation matrix. Briefly, the soil sample is lysed using Lysis Solution in combination with the provided Bead Tubes. The released RNA is then bound to Norgen's column in the presence of Binding Solutions (BIND). Under these conditions only the RNA will bind to the column, while the contaminating proteins will be removed in the flowthrough or retained on the top of the resin. The bound RNA is then washed to remove any remaining impurities (WASH). Lastly, the purified inhibitor-free RNA is eluted into the provided Elution Buffer (ELUTE). Please see procedure flowchart below.
Norgen’s Soil Total RNA Purification Kit purifies all sizes of RNA, from large mRNA and ribosomal RNA down to microRNA and small interfering RNA. The protocol does not rely on the use of phenol or chloroform, thereby providing a user friendly procedure and allowing high-throughput analysis on the lab bench. The purified RNA is of the highest integrity, and can be used in a number of downstream applications including real time PCR and reverse transcription PCR for gene expression analysis.
Performance Data

Fig.1. Isolation of Total RNA from Bacteria in Soil. Pseudomonas fluorescens was spiked into 250 mg samples of autoclaved soil and total RNA was isolated using Norgen’s Soil Total RNA Purification Kit. RNA was visualized by running 7.5 µL of each 75 µL elution on a 1.2% agarose-formaldehyde RNA gel. Total RNA (large and small) of Pseudomonas fluorescens was recovered from the autoclaved spiked soil without any significant degradation, indicating that high integrity RNA can be purified from the microorganisms in the soil. Lanes 1 and 2 contain total RNA from Pseudomonas fluorescens, Lanes 3 and 4 contain total RNA purified from the autoclaved soil spiked with Pseudomonas fluorescens, and Lanes 5 and 6 contain RNA purified from the autoclaved soil (no RNA was found).
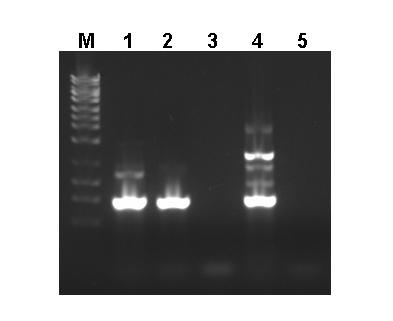
Fig.2. High Quality RNA Free from PCR Inhibitors. Pseudomonas fluorescens was spiked into 250 mg samples of autoclaved soil and total RNA was isolated using Norgen’s Soil Total RNA Purification Kit. One microliter of each elution was then used as the template in a 20 µL RT-PCR reaction to detect the 16s rRNA. Lane 1 contains the results when total RNA from Pseudomonas fluorescens was used as the input, Lane 2 is the results when total RNA isolated from the autoclaved soil spiked with Pseudomonas fluorescens was used as the input, Lane 3 is the results when RNA purified from non-spiked autoclaved soil was the input, Lane 4 is a positive control and Lane 5 is the Negative control. As it can be seen the RNA purified from soil using Norgen's kit was of a high quality and can be successfully used in sensitive downstream applications.
Product Citations
- Giovambattista Sorrenti, Giampaolo Buriani, Francesca Gaggìa, Loredana Baffoni, Francesco Spinelli, Diana Di Gioia, Moreno Toselli (2017) Soil CO2 emission partitioning, bacterial community profile and gene expression of Nitrosomonas spp. and Nitrobacter spp. of a sandy soil amended with biochar and compost Applied Soil Ecology
- Maria De Angelis, Eustacchio Montemurno, Maria Piccolo, Lucia Vannini, Gabriella Lauriero, Valentina Maranzano, Giorgia Gozzi, Diana Serrazanetti, Giuseppe Dalfino, Marco Gobbetti, Loreto Gesualdo. (2014) Microbiota and Metabolome Associated with Immunoglobulin A Nephropathy (IgAN). Microbiota and Metabolome of IgA Nephropathy Patients
- Wang Y, Hayatsu M, Fujii T. (2012) Extraction of Bacterial RNA from Soil: Challenges and Solutions. Microbes and the Environment / JSME
- Liang Z, Keeley A. (2011) Detection of viable Cryptosporidium parvum in soil by reverse transcription real time PCR targeting hsp70 mRNA. Applied and Environmental Biology
- Catalog Number
27750-NB - Supplier
Norgen Biotek - Size
- Shipping
RT
You save 10 %
578,00 €
