Total RNA Purification Kit (96-well column chromatography)
| Specifications | |
|---|---|
| Product Category: | Total RNA incl. microRNA Purification |
| Sample Type: | Cultured animal cells, tissue samples, blood, plasma,serum, bacteria, yeast, fungi, plants |
Product Description
- Isolate total RNA, from large rRNA down to microRNA (miRNA)
- Isolate high quality total RNA with complete size range from a variety of sources, even from plasma and serum
- Isolate viral RNA from swab samples
- RNA can be isolated and detected from as little as a single animal cell (see application note - link below)
- No phenol or chloroform extractions
- Fast and easy processing (Lack of phase extraction step allows for easy automation/high throughput applications)
- 96-well plates can be rapidly processed in 30 minutes with vacuum manifold or centrifuge. No need for special equipment.
- Kits with spin columns for individual preparations also available
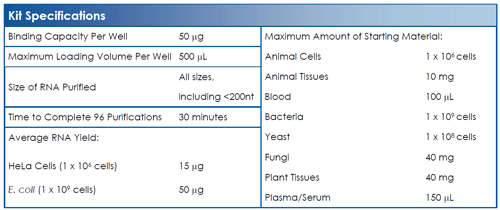
Norgen’s Total RNA Purification 96-Well Kit provides a rapid method for the high-throughput isolation and purification of total RNA using a 96-well format in 30 minutes. The kit purifies all sizes of RNA, from large mRNA and ribosomal RNA down to microRNA (miRNA) and small interfering RNA (siRNA) without the use of inhibitory phenol or chloroform. Total RNA can be isolated from a broad range of sample sources including cultured cells, tissues, blood, serum, plasma, bacteria, yeast, fungi, plants and viruses. The kit is highly sensitive, as RNA can be isolated from as little as a single cell.
Purification Technology
Purification is based on 96-well spin column chromatography using Norgen’s proprietary Silicon Carbide (SiC) resin as the separation matrix. Purification can be performed on either a vacuum manifold or using centrifugation. Briefly, the cells or tissue of interest is first lysed using the provided Lysis Solution, ethanol is added and the RNA is bound to Norgen's 96-well filter plates . Under these conditions only the RNA will bind to Norgen's resin while most of the contaminating cellular proteins are removed in the flowthrough or retained on top of the resin. The bound RNA is then washed to remove any remaining impurities. Lastly, the purified total RNA is eluted into 75 uL of the provided Elution Buffer or water.
Norgen’s proprietary Silicon Carbide resin provides superior affinity to the full size range of RNA molecules, resulting in large and small RNA (miRNA) purification with excellent consistency and sensitivity. The purified RNA is of the highest integrity, and can be used in a number of downstream applications including real time PCR, reverse transcription PCR, Northern blotting, RNase protection and primer extension, expression array assays, miRNA cloning and Next Generation Sequencing.
High Quality of Total RNA from a Diverse Range of Sample Sources
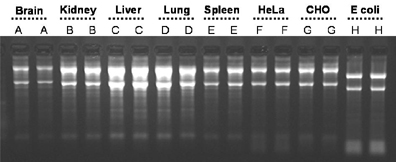
Norgen’s Total RNA Purification 96-Well Kit allows for RNA isolation from a wide range of species and tissue types. Total RNA was isolated from 8 mg of brain (Lanes A), kidney (Lanes B), liver (Lanes C), lung (Lanes D) and spleen tissue (Lanes E), 7.5 x 10e5 HeLa (Lanes F) and CHO cells (Lanes G), and 5 x 10e8 bacteria (Lanes H) using Norgen’s Total RNA Purification 96-Well Kit. Five microliters of the 75 uL isolated RNA was resolved on a 1.2% formaldehyde agarose gel. It can been seen that Norgen’s kit can be used to successfully isolate total RNA, including small RNA species, from a broad range of sample types.
High Quality of Total RNA with Complete Size Range
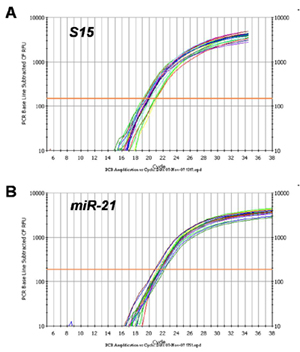
Consistent Isolation of Total RNA, including microRNA:
Total RNA was isolated from samples of 5 x 10^5 HeLa cells using Norgen’s Total RNA Purification 96-Well Kit. Aliquots of each total RNA sample were then used in 2 different RT-qPCR reaction. In the first RT-qPCR the S15 gene of mRNA was amplified (Panel A), and in the second RT-qPCR reaction the miR-21 microRNA was amplified (Panel B). Both the mRNA and microRNA were amplified in a consistent manner from all the samples, with low variability of the C T values. Thus both types of RNA are being consistently isolated using this kit.
Great Isolation Sensitivity
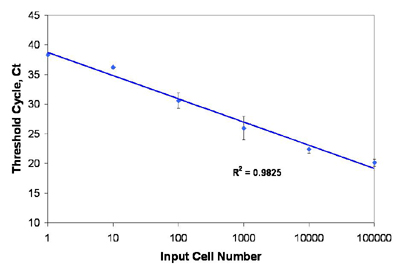
Norgen’s Total RNA Purification Kit allows sensitive RNA extraction from even less than 10 cells. RT-qPCR was used to detect mRNA isolated from various input amounts of HeLa cells, from 100,000 down to a single cell, using Norgen’s Total RNA Purification 96-Well Kit. Ten microliters of the 75 µL isolated RNA was then subjected to a 20 µL reverse transcription using oligo dT primer. Three microliter of the reverse transcription was used in a 20 µL real-time PCR reaction with primers to detect the human S15 transcripts. The resulting Ct values were plotted against the input cell number. Total RNA was isolated and detected linearly from as little as a single HeLa cell.
Typical RNA Yields
HeLa Cells (1 x 10e6 Cells): 15 ug
E coli (1 x 10e9 Cells): 50 ug
Brain (10 mg): 9 ug
Liver (10 mg): 26 ug
Blood (50 uL, Hamster) 1 ug
Total RNA Purification 96-Well Kit Applications
- Quantitative, real-time RT-PCR for both large mRNA and small RNA including miRNA
- RT-PCR for both large mRNA and small RNA including miRNA
- Expression Profiling
- Next Generation Sequencing for RNA and miRNA
- miRNA, also from plasma and serum, for discovery
- microRNA cloning and amplification
- PCR-based virus detection
- PCR-based viable bacteria detection
- Northern Blotting
- RNase Protection
- Primer Extension
Product Citations
- Mestdagh P, Gillard M, Dhillon SK, et al. (2021) Evaluating Diagnostic Accuracy of Saliva Sampling Methods for Severe Acute Respiratory Syndrome Coronavirus 2 Reveals Differential Sensitivity and Association with Viral Load The Journal of Molecular Diagnostics.
- Babouee Flury B, Ellington MJ, Hopkins KL, Turton JF, Doumith M, Loy R, Staves P, Hinic V, Frei R, Woodford N (2016) Association of novel nonsynonymous single nucleotide polymorphisms in ampD with cephalosporin resistance and phylogenetic variations in ampC, ampR, ompF and ompC in Enterobacter cloacae that are highly resistant to carbapenems Antimicrobial Agents and Chemotherpy
- Fishilevich E, Vélez AM, Khajuria C, Frey ML1, Hamm RL, Wang H, Schulenberg GA, Bowling AJ, Pence HE, Gandra P, Arora K, Storer NP, Narva KE, Siegfried BD (2016) Use of chromatin remodeling ATPases as RNAi targets for parental control of western corn rootworm (Diabrotica virgifera virgifera) and Neotropical brown stink bug (Euschistus heros) Insect Biochemistry and Molecular Biology
- Michelle J. Ormseth, Joseph F. Solus, Kasey C. Vickers, Annette M. Oeser, Paolo Raggi and C. Michael Stein (2015) Utility of Select Plasma MicroRNA for Disease and Cardiovascular Risk Assessment in Patients with Rheumatoid Arthritis The Journal of Rheumatology
- Li W, Turner A, Aggarwal P, Matter A, Storvick E, Arnett DK, Broeckel U (2015) Comprehensive evaluation of AmpliSeq transcriptome, a novel targeted whole transcriptome RNA sequencing methodology for global gene expression analysis. BMC Genomics
- Catalog Number
24300-NB - Supplier
Norgen Biotek - Size
- Shipping
RT
You save 10 %
856,00 €
