
CRISPR-based NGS Library Depletion
CRISPR-based NGS Library Depletion
Don’t Waste your Valuable Reads for Repeatedly Captured Sequences
The ability to sequence millions of reads from genomes and transcriptomes has yielded an unprecedented discovery on genes and genetic variation. Unfortunately, much of these reads are wasted on over abundant sequences that are captured repeatedly.

Future discovery may be accelerated by cutting through the noise of over abundant sequences. Shifting focus away from abundant genes and increasing sensitivity by lowering background noise is required to enable this discovery. It’s why our partner Jumpcode created DepleteX.
DepleteX Technology from Jumpcode Genomics harnesses the specificity of CRISPR-Cas9 to degrade abundant, uninformative sequences in next-generation sequencing (NGS) libraries. With DepleteX, you can see more lower expressing transcripts and shift your sequencing power for deeper coverage and improved signal.
Now you can see exactly what you want, breaking through the clutter - and breaking new ground.
How DepleteX works
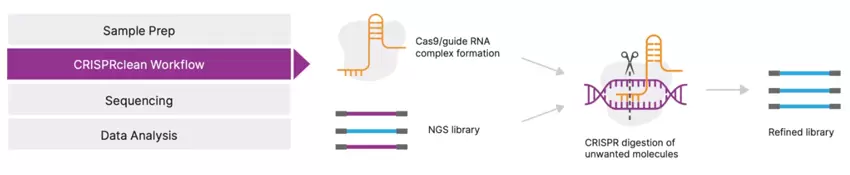
DepleteX-mediated depletion can be integrated directly into NGS library prep protocols at various points depending on the application. CRISPR-Cas9 complexes are formed with a pool of designed guide RNAs. After targeted sequences are cut, they cannot be a substrate for PCR amplification or subsequent sequencing. The output is a refined, sequence-ready NGS library.
Benefits
- Highly programmable
- Increased sensitivity
- Simple automatable workflow
- Genomic analysis platform agnostic
Applications
DepleteX technology is both specific and highly programmable to remove unwanted sequences for many applications. View the applications below to learn how Jumpcode is driving discovery today.
Whole Transcriptome Profiling
~90% of total RNA is abundant ribosomal RNA noise. This high abundance of ribosomal sequences in samples often results in more background noise than usable data. Detecting novel transcripts or low-expressing transcripts becomes a time-consuming challenge, where the biologically relevant data you need is often lost in the clutter. With CRISPRclean Ribodepletion Kits, you can access an unbiased whole transcriptome – without the noise.
Single Cell Analysis
Boost usable single cell RNA seq data and gain a deeper view of expression profiles of individual cells with the DepleteX Single Cell RNA Boost Kit. The kit depletes sequences not used for secondary analysis including non-transcriptomic reads, ribosomal RNA, ribosomal and mitochondrial mRNA as well as non-variable genes (optional).
Microbiome
Improve microbial genome coverage and increase detection sensitivity of low abundant microbial species with DepleteX. Human microbiome sample types are diverse and come from a variety of body sites including saliva, gut, skin, wound infections and everywhere a microbial community can be found. Samples often consist of a mixture of human and microbial cells. For some samples like stool, the composition is generally made up of mostly bacterial species and very little human cells, but nasal swabs is the opposite. One commonly used method for human microbiome research is shotgun metagenomics sequencing, which is used to characterize the complete bacterial phylogeny and taxonomy of complex, microbiome samples. Depending on the sample type and the research goal, human-derived complex samples can be a challenge due to the overwhelming human host contamination that obscures sequencing coverage of interesting microbial genomes.
With the DepleteX Human DNA Depletion Kit or the CRISPRclean Plus Stranded Total RNA Prep with rRNA Depletion Kit you can gain an accurate taxonomic profile of a microbial community by removing human host DNA or RNA for shotgun metagenomics or metatranscriptomics.
Infectious Disease Surveillance
Detect known and novel pathogens to accelerate outbreak investigations. You can acquire viral genomic data, microbiome composition, information on co-infections, and host-immune response in a single workflow with the CRISPRclean Plus Stranded Total RNA Prep with rRNA Depletion. Please refer to the Application Note CRISPRclean Plus for SARS-CoV-2 Shotgun Metatranscriptomic Sequencing for further information.
Custom depletion panels for your samples and applications are available upon request, please contact us if you are interested.
DepleteX Depletion Panels
In addition to the library preparation kits we also offer Jumpcode's DepleteX Depletion Panels with different focused panels for increasing detection sensitivity of low expressing transcripts and one broad RNA depletion panel to maximize low expressing transcripts.
Please note: CRISPRclean is now DepleteX